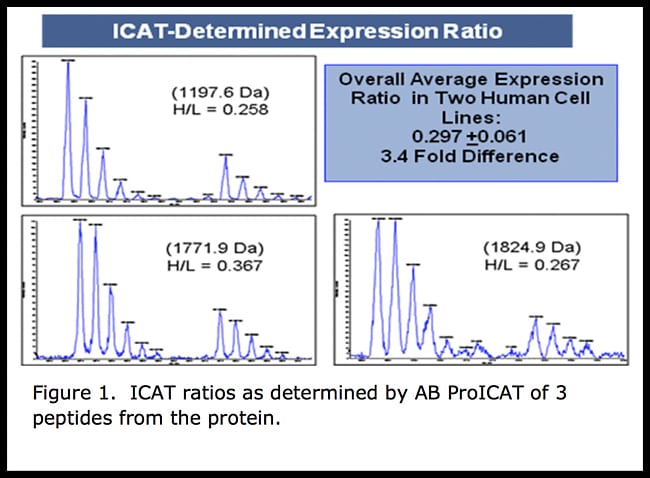
The Application of New Software Tools to Quantitative Protein Profiling Via Isotope-coded Affinity Tag (ICAT) and Tandem Mass Spectrometry - Molecular & Cellular Proteomics

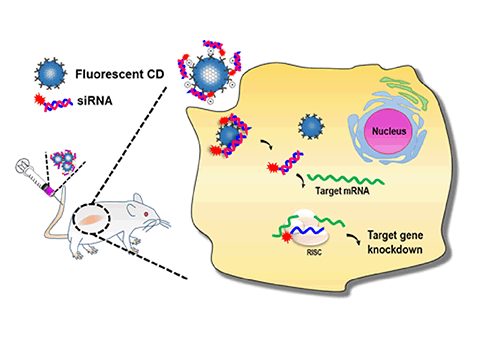
Simultaneous Identification and Quantification of Nitrosylation Sites by Combination of Biotin Switch and ICAT Labeling | SpringerLink

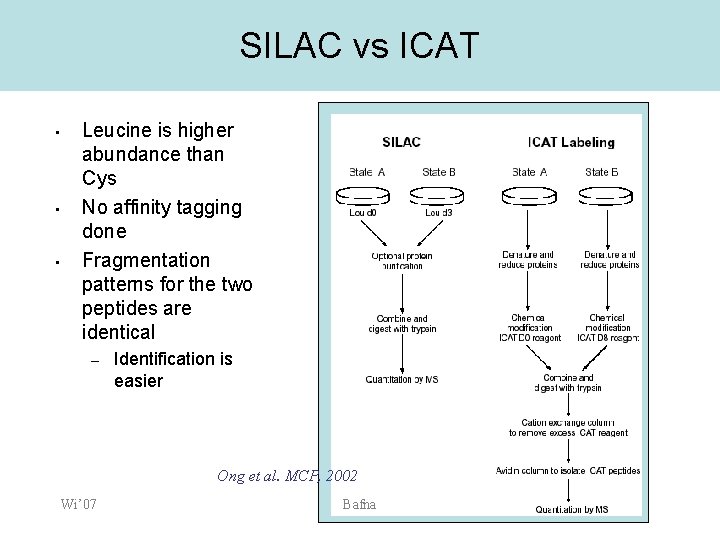
Quantitative Mass Spectrometry-Based Approaches in Cardiovascular Research | Circulation: Cardiovascular Genetics

Figure 1 from The use of isotope-coded affinity tags (ICAT) to study organelle proteomes in Arabidopsis thaliana. | Semantic Scholar
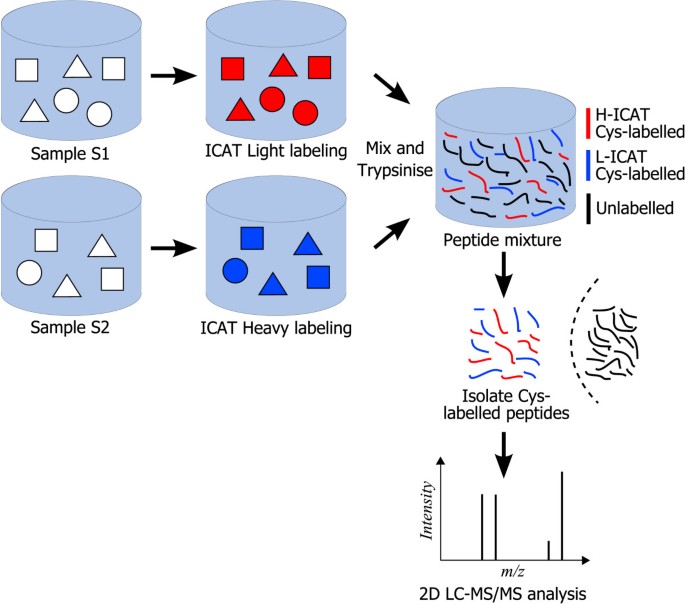
The EIPeptiDi tool: enhancing peptide discovery in ICAT-based LC MS/MS experiments | BMC Bioinformatics | Full Text

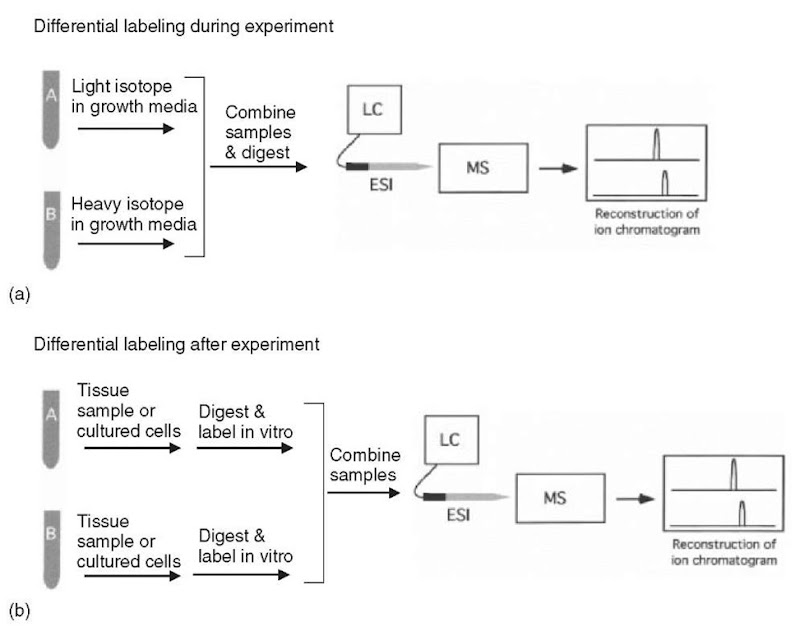
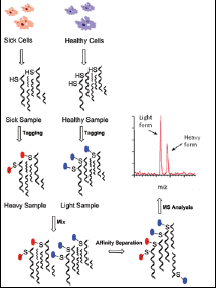
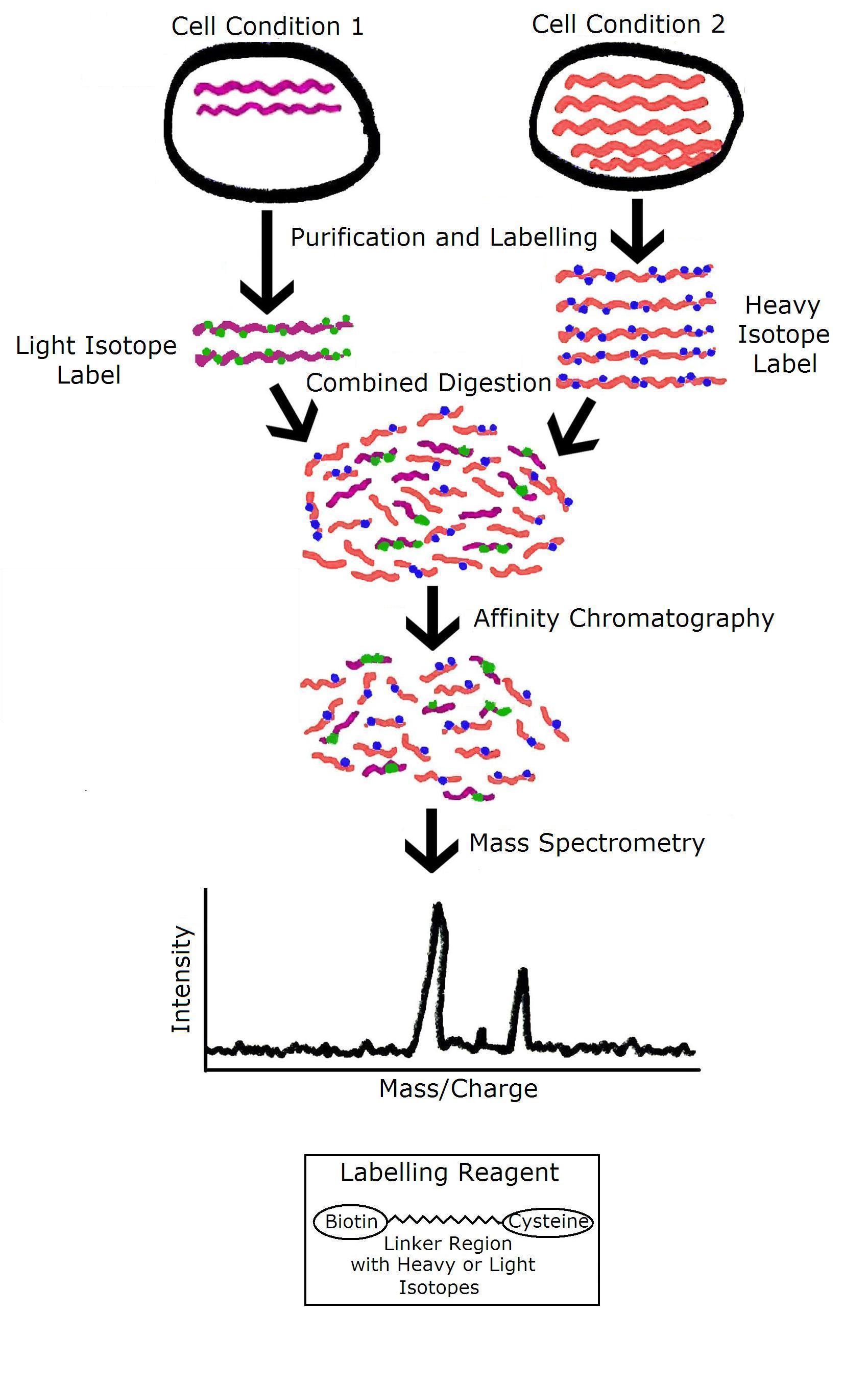
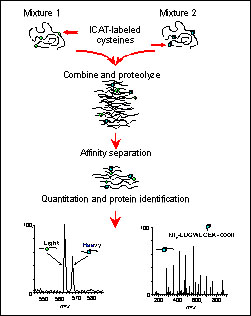
A) ICAT reagent and (B) strategy for quantitation by ICAT. Two protein... | Download Scientific Diagram

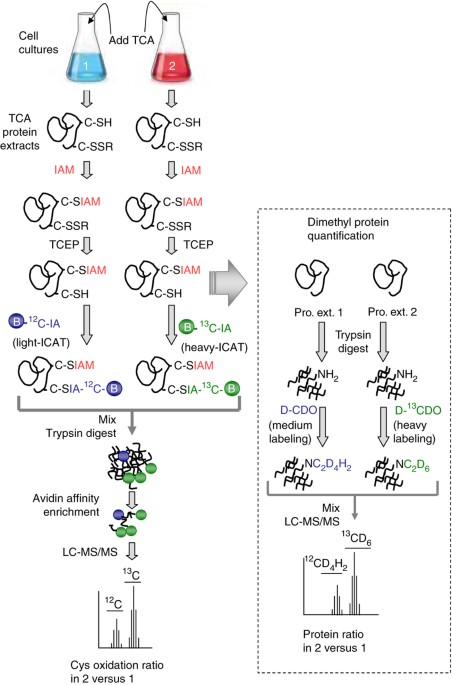
Isotope-coded Affinity Tag Approach to Identify and Quantify Oxidant-sensitive Protein Thiols - ScienceDirect

An Optimized Strategy for ICAT Quantification of Membrane Proteins* - Molecular & Cellular Proteomics