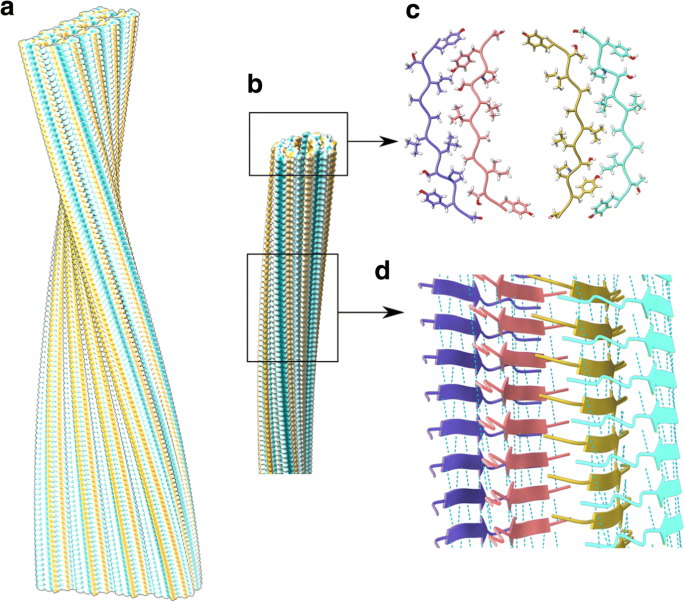
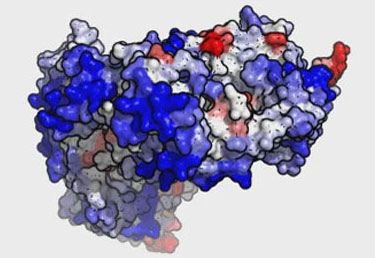
Cryo-EM structures of amyloid fibrils from recombinant SAA1.1 protein... | Download Scientific Diagram

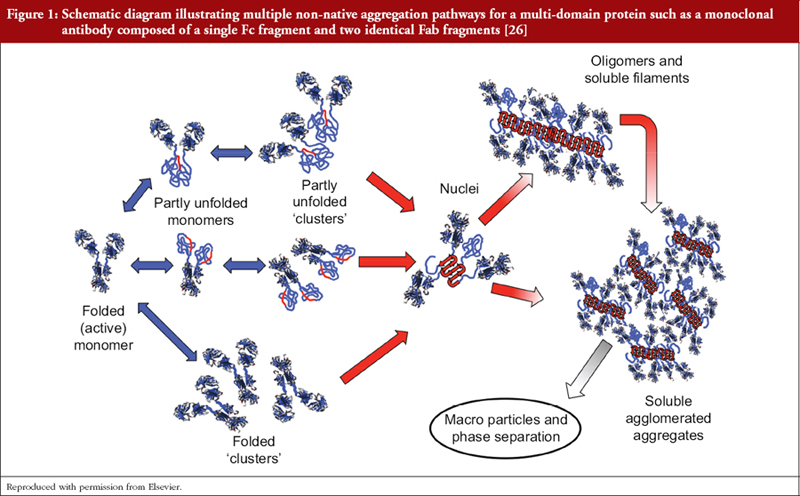
Using extensional flow to reveal diverse aggregation landscapes for three IgG1 molecules - Willis - 2018 - Biotechnology and Bioengineering - Wiley Online Library

Computational prediction of protein aggregation: Advances in proteomics, conformation-specific algorithms and biotechnological applications - ScienceDirect

The signal peptide of the amyloid precursor protein forms amyloid-like aggregates and enhances Aβ42 aggregation - ScienceDirect

Electrostatic interactions drive native‐like aggregation of human alanine:glyoxylate aminostransferase - Dindo - 2017 - The FEBS Journal - Wiley Online Library

Computational prediction of protein aggregation: Advances in proteomics, conformation-specific algorithms and biotechnological applications - ScienceDirect

Structure-aware protein solubility prediction from sequence through graph convolutional network and predicted contact map | Journal of Cheminformatics | Full Text

Computational prediction of protein aggregation: Advances in proteomics, conformation-specific algorithms and biotechnological applications | Semantic Scholar
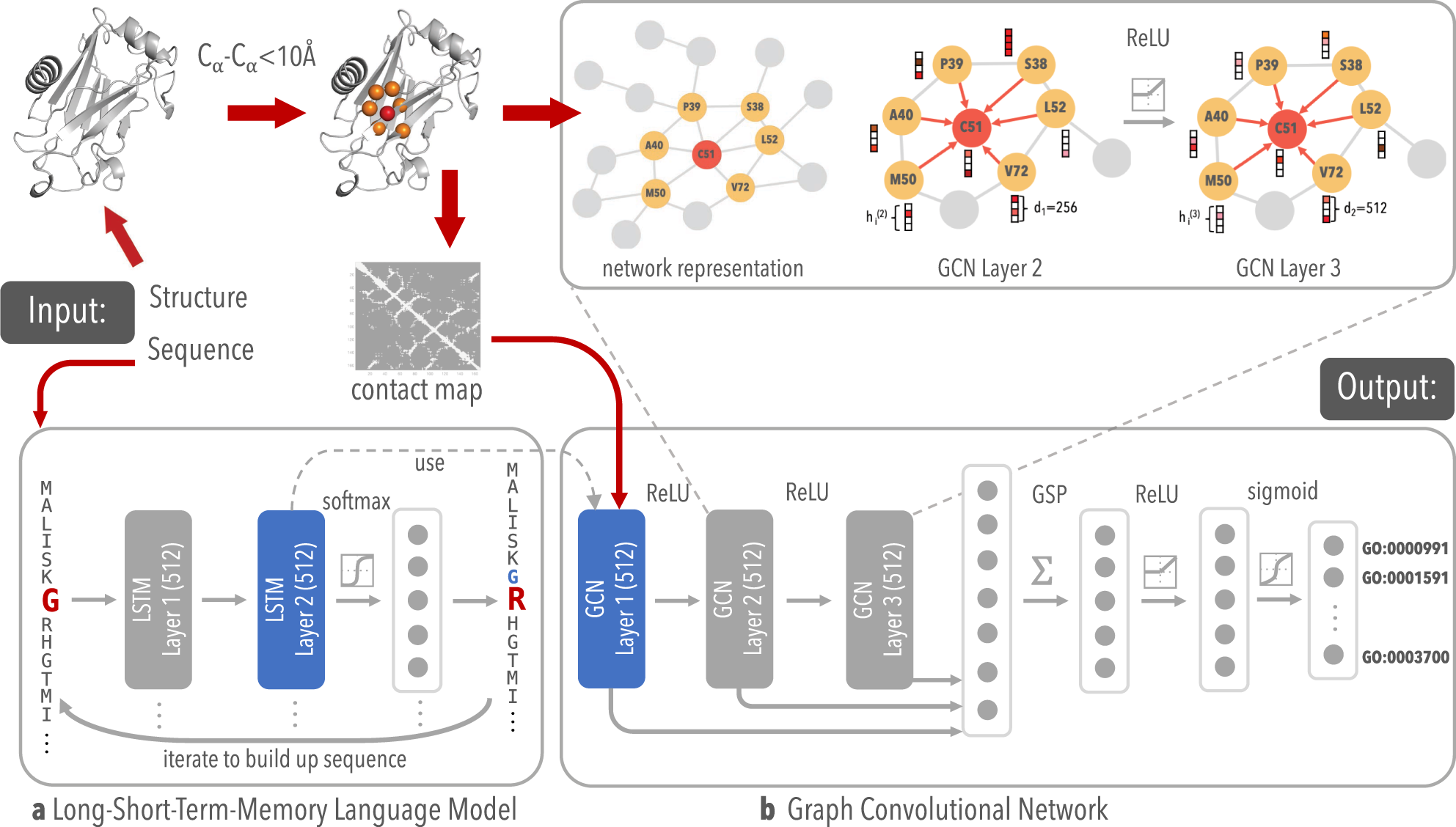
Structure-based protein function prediction using graph convolutional networks | Nature Communications
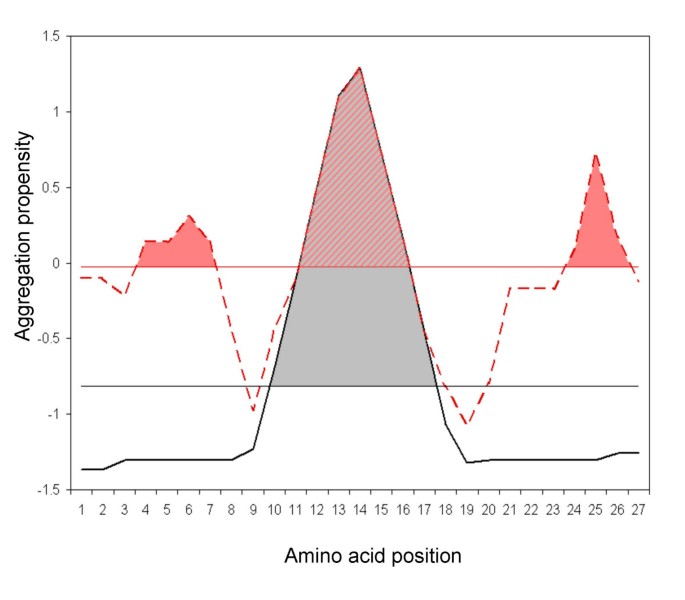
AGGRESCAN: a server for the prediction and evaluation of "hot spots" of aggregation in polypeptides | BMC Bioinformatics | Full Text
PLOS ONE: Genome-Wide Prediction and Analysis of 3D-Domain Swapped Proteins in the Human Genome from Sequence Information